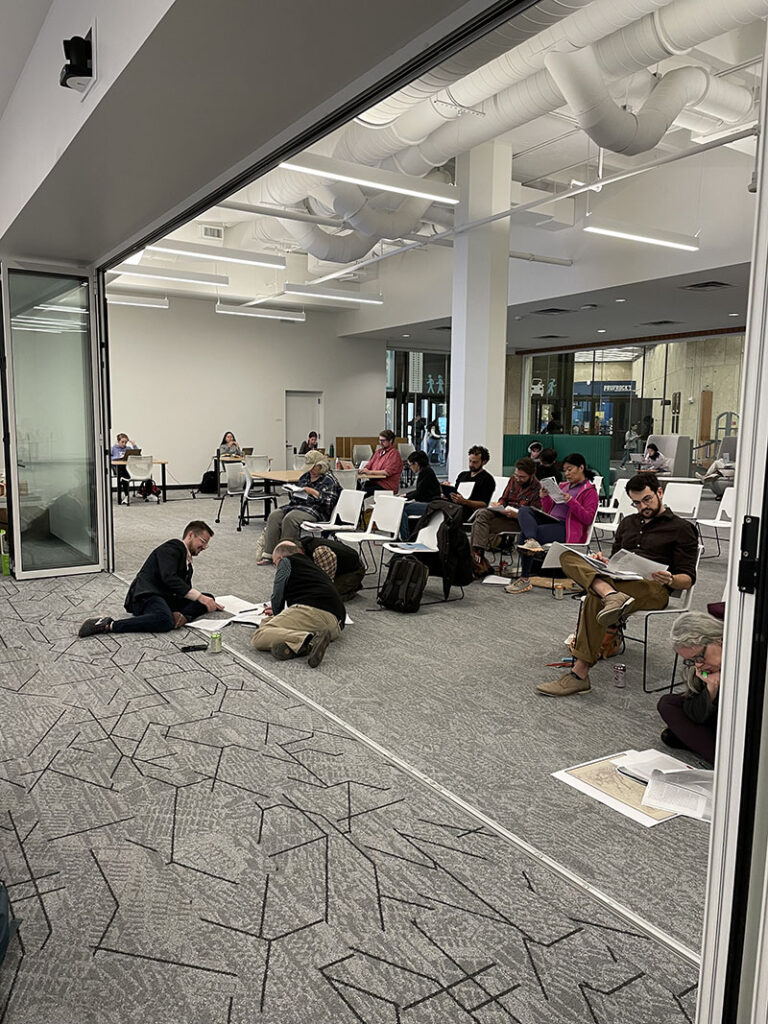
The Librarian Lens is an occasional column featuring librarians who support the research lifecycle across a range of disciplines sharing research tips, updates about both Libraries-provided and open source resources, and related topics intended to intrigue, demystify and highlight topics of interest to the research-curious. Posted columns are provided or curated by librarians from the University of Texas Libraries STEM and Social Science Engagement Team.
Calling all researchers, students and anyone interested in exciting developments in health and medical information…read on to find out about the exciting All of Us Research Program that is on a mission to accelerate medical breakthroughs!
The All of Us Research Program is part of the National Institutes of Health. It is actively collecting data from a diverse population of participants across the United States. As of April 1, 2024 the program has 783,000+ participants, 432,000+ electronic health records and 555,000+ biosamples received according to the program’s Data Snapshots, which are updated daily. The program continues to enroll people from all backgrounds and is aiming for 1,000,000+ participants.
The Data
The program aims to connect the research community with the data, which is longitudinal and from a diverse population, including those typically underrepresented in biomedical research. The data are from participants’ survey responses, electronic health records, physical measurements, Fitbit records and biosamples. DNA from participants’ biosamples supply genomic data, including genotyping arrays and whole genome sequences. The data undergoes a curation process with rigorous privacy protections. On average the data is pushed out once a year. The All of Us Research Hub stores the data in a secure Cloud-based space, where it must remain. Imagine the possibility of all the research project opportunities, especially those on precision medicine, with the ability to combine genomic data with phenotypic data and analyze it on a large scale!
Public Access
There is a terrific way to get a better idea of the data that are available whether out of curiosity or for more in-depth study and that is through the program’s Data Snapshots and interactive Data Browser that anyone can access through the All of Us Research Hub. No prior authorization or account is required to explore the aggregated, anonymized data. In fact, this public access data can serve as a great jumping off point for developing a research project question! In addition, the Research Projects Directory, which currently lists over 10,000 projects, can be viewed by anyone and is a great place to browse for inspiration or collaboration opportunities. For students, it is a wonderful way to see examples of scientific questions and approaches. Papers resulting from these research projects are listed on All of Us Publications and anyone can read them because they have been published open access. The total number of publications is close to 300!

Credit: Jonathan Bailey, NHGRI. Public Domain
Researcher Access
The All of Us Research Hub provides access to more granular data along with tools for analysis through the Researcher Workbench. To access granular data either on the registered tier or the controlled tier, researchers must complete registration and be approved. Registration is a four step process. The first is to confirm your institution has a Data Use and Registration Agreement (DURA) in place with All of Us. At the time of this writing, the University of Texas at Austin has a DURA in place. The next step is to create an account and verify your identity. The third is to complete mandatory training. Additional training is required to analyze the controlled tier data because that is where the genomic data is stored. The fourth and final step is to sign the Data User Code of Conduct. Steps two through four take about two hours to complete but do not have to be done all in one sitting. After completing registration, researchers will be able to access the data once they receive notification of approval from the All of Us program. The data can then be accessed and analyzed through the Researcher Workbench where researchers create Workspaces and submit descriptions of their research projects for listing on the Research Projects Directory. The program makes it easy for researchers to invite other registered users from their own institutions or from other institutions to their Workspaces for collaboration. This is especially helpful if a researcher needs someone else to run analysis with R or Python, the two options currently available. The program encourages and supports team science!
Access Cost & User Support
Even though there is a cost to researchers for analyzing and storing data for their research projects in the Cloud, there is good news. The program provides researchers $300 initial credits upon creating Workspaces in the Researcher Workbench to help them get started on their research projects. Researchers can get information on how to run their data cost effectively from the program’s User Support Hub. They can also speak with the program’s support team to get an idea of cost before analyzing data.
The All of Us Research Program’s investment in medical breakthroughs could not be more clear with the amount of thoughtful, varied support it has put together to help researchers access and analyze the data. The User Support Hub has video tutorials, articles, a help button, and an event calendar that lists the support team’s upcoming office hours among other events. Support resources for the Researcher Workbench range from getting started to working with the data to credits and billing. The support articles are kept current and are on topics such as using RStudio on the Researcher Workbench. The support team is responsive and ready to help! Help is provided by actual people, not chatbots. The program welcomes feedback and uses it to make the program better. For instance, if there is certain data not being collected that would be helpful to collect, let the program know!

Credit: Darryl Leja, NHGRI. Public Domain
The Impact
By collecting data from the United States’ diverse population, especially from traditionally underrepresented groups in biomedical research, the All of Us Research Program strives to make the medical data that is currently available to researchers more complete. With researchers having access to not only genomic data, but also phenotypic data, their studies can fuel insights into our health on an individual level; thus potentially allowing for healthcare providers to provide better, more tailored care to each of us. To see the research studies made possible so far by the availability of the All of Us data, take a look at the All of Us Research Highlights today, and check back regularly to keep an eye on advancements and breakthroughs!
The All of Us Research Program seeks to connect the research community with its dataset, which is one of the largest, most diverse ever assembled, to spur research that will improve health for all of us. If you are feeling inspired as a researcher or want to become a participant or are just curious to find out more about the program, visit the program’s websites at researchallofus.org or allofus.nih.gov.